
Andrew P. Spicer
Andrew Spicer grew up in both England and the United States. He has always had a passion for biology, ranging from hobbies that include insect rearing and collecting and ornithology to his formal research interests in glycobiology and mouse genetics. He received his Ph.D. from the Imperial Cancer Research Fund and University College in London in 1993. His Ph.D. thesis work focused upon studies of an epithelial mucin, Muc1, in the mouse. In 1992, he moved to the United States, where he helped establish the laboratory of Dr. Sandra Gendler, his Ph.D. supervisor, at Mayo Clinic Scottsdale in Arizona. In 1994 he became a Postdoctoral Fellow in Dr. John McDonald's laboratory there. His postdoctoral research led to an important breakthrough in hyaluronan biology--the identification of a family of genes encoding vertebrate hyaluronan synthases. In August 1997 he established an independent laboratory as Assistant Professor within the Rowe Program in Genetics and the Department of Biological Chemistry in the School of Medicine at the University of California, Davis. His laboratory continues to study the vertebrate hyaluronan synthases, using the mouse as a model to investigate the role of hyaluronan and other glycosaminoglycans in embryonic development and in adult tissues.

John A. McDonald
John McDonald studied at the University of South Florida, where the late Dr. Jerome O. Krivanek, Chairman of Zoology, was a pivotal influence. Dr. McDonald received a Ph.D. in Biochemistry from Rice University in 1970 and the M.D. from Duke University School of Medicine in 1973. After training at the Peter Bent Brigham Hospital in Boston, Massachusetts, and the National Institutes of Health in Bethesda, Maryland, he joined the faculty at Washington University Medical School (St. Louis, Missouri) in 1979. From 1985 until 1991, he was professor of medicine and cell biology and Director of the Respiratory and Critical Care Division at Barnes Hospital-Washington University Medical Center, leading a significant expansion of training programs and research efforts. In 1991 he joined the Mayo Clinic Scottsdale, Arizona, as Director for Research, orchestrating a major expansion of basic and clinical research. Dr. McDonald's principal laboratory was the first to elucidate a critical role for integrin cell surface receptors in extracellular matrix assembly and to define a high-resolution biological map of fibronectin. He has also contributed to our understanding of architectural remodeling in human lung disease and the role of matrix receptors in lung development and cardiac remodeling. Recently, his laboratory began to investigate the eukaryotic family of hyaluronan synthases, the topic of this overview.
Hyaluronan is a polymer containing multiple copies of the disaccharide N-acetyl-D-glucosamine (GlcNAc) and D-glucuronate (GlcA). Most glycosaminoglycans are synthesized in the Golgi network within the cell and are attached to proteins. In contrast, hyaluronan is synthesized at the plasma membrane and is not associated with any core protein (see references 1 and 2 for review). Biochemical and cell biological studies have revealed that hyaluronan is synthesized at the inner face of the cell membrane. The growing polymer is extruded through the membrane to the outside of the cell as it is being synthesized (Fig. 1).3 Outside of the cell, hyaluronan can form a complex with hyaluronan-binding proteins (including members of the lectican family, such as aggrecan or versican) and with inter-alpha trypsin inhibitor to form a pericellular coat. This coat can be visualized around mammalian cells growing in tissue culture by the addition of small particles, such as fixed red blood cells, to the tissue culture medium. Upon settling, these particles are excluded from the cell membrane by the large size and negative charge of the hyaluronan coat (Fig. 1). It has been suggested that the assembly and presence of such a hyaluronan-dependent pericellular coat may be required for migration and/or proliferation of certain cell types, such as vascular smooth muscle cells.a
a Topic of future article by Tom Wight.
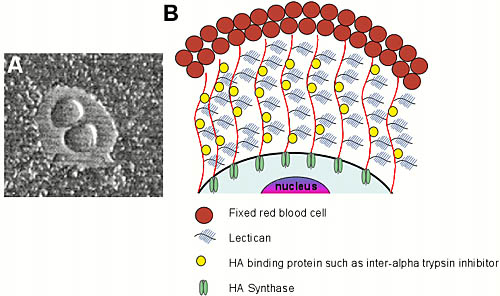
Fig. 1 Hyaluronan biosynthesis occurs at the plasma membrane in eukaryotic cells.
(a) Hyaluronan-dependent pericellular coat, as revealed by a particle exclusion assay. Fixed sheep erythrocytes were added to the culture dish and allowed to settle on the monolayer of cells. The erythrocytes are excluded from the cell membrane by the large size and charge of the pericellular coat. The coat can be formed in two ways: cells that actively synthesize hyaluronan extrude the newly synthesized polymer through the plasma membrane, where it can form a complex with hyaluronan-binding proteins. Alternatively, cells that do not actively synthesize hyaluronan but that possess high-affinity cell surface hyaluronan receptors can assemble a pericellular coat with exogenous hyaluronan and hyaluronan-binding proteoglycans.
(b) Cartoon of hyaluronan biosynthesis and pericellular coat formation in vertebrate cells. In this cartoon, only cells that form a pericellular coat in a hyaluronan synthase-dependent fashion are considered.
Synthesis of hyaluronan at the plasma membrane suggests that the required enzymatic machinery would be located within or at the cell membrane. Accordingly, attempts were made to solubilize and partially purify the mammalian hyaluronan synthase from the plasma membrane of hyaluronan-synthesizing cells. However, little progress was made toward the identification of the enzymes involved in hyaluronan biosynthesis until 1996, when several laboratories independently reported the identification of new genes encoding potential hyaluronan synthases (reviewed in Ref. 2). In this article, we review the advances that have been made in this rapidly moving field since the molecular cloning of the first putative eukaryotic hyaluronan synthases.
A Note on Nomenclature of Hyaluronan Synthases.In Group A streptococci, such as Streptococcus pyogenes, polymerization of hyaluronan from its activated UDP-sugars occurs through the action of a single protein, hasA or spHAS.3 Furthermore, the purified hasA protein possesses hyaluronan synthase activity in vitro.4 Thus, a single protein can catalyze the formation of two distinct glycosidic bonds, the beta1,3 and beta1,4 bonds that link glucuronate to N-acetylglucosamine and N-acetylglucosamine to D-glucuronate. This capability goes against the classical dogma of one gene, one protein, and one function. Not only is there compelling evidence that this single protein can polymerize hyaluronan from its precursors, but there is also evidence to suggest that this protein may be involved in the translocation of the growing polymer across the cell membrane.
Sequence analyses indicated that the primary amino acid sequence for hasA shared some identity with other beta-glycosaminyltransferases, including the nodulation factor (NodC) enzyme of root-nodule-colonizing Rhizobium species and the chitin synthases of Saccharomyces cerevisiae and other fungi. In addition, these enzymes shared some identity with a previously identified cDNA, designated DG42, in the vertebrate developmental model organism Xenopus laevis (the African clawed toad). We, and other investigators, capitalized upon these sequence identities to design degenerate PCR strategies that led to the identification and isolation of cDNAs encoding putative vertebrate hyaluronan synthases. cDNAs encoding two related putative hyaluronan synthases, Has2 and Has3, were identified in human, mouse, chicken, Xenopus laevis, and the zebrafish.5, b The sequences were highly conserved between different species. An alternative expression cloning approach led to the identification of an additional putative vertebrate hyaluronan synthase, designated Has1, in mouse and human. cDNA clone pools were transfected into cells deficient in the ability to synthesize hyaluronan, and the transfected cells were screened to identify those cells that had acquired hyaluronan biosynthetic activity. Most recently, we have identified the first invertebrate hyaluronan synthase genes, in the chordate Amphioxus,b demonstrating that hyaluronan biosynthesis predates vertebrate evolution. It is possible that the ability to synthesize hyaluronan evolved alongside the ability to form a more advanced body plan, including a notochord.
b Spicer, A.P., manuscript in preparation.
Sequence analyses predict that all eukaryotic hyaluronan synthases and the streptococcal hyaluronan synthase encode plasma membrane proteins with multiple transmembrane domains. There are two predicted amino-terminal transmembrane domains and a cluster of C-terminal transmembrane domains, separated by a relatively large intracellular (cytoplasmic) domain, predicted to contain sequences responsible for the catalytic activities of these proteins.3 This structure is very similar to the predicted structures for related beta-glycosaminyltransferases, such as NodC and the cellulose synthases (CelAs) of plants. Furthermore, sequence alignments have identified invariant residues within the predicted intracellular loop of all of these proteins,5 suggesting that these residues are required for catalytic activity. Is it conceivable that the hyaluronan synthase, chitin synthase, and cellulose synthase genes of eukaryotes share a common ancestor and have evolved through divergent evolution? In humans and mice, there are three hyaluronan synthases, encoded by related yet distinct genes located on different chromosomes. The encoded proteins share 55-71% amino acid sequence identity (Fig. 2).

Fig. 2 Multiple amino acid sequence alignment of vertebrate hyaluronan synthases.
Alignment of mouse (m), human (h), and Xenopus laevis (x) Has1, Has2 and Has3 amino acid sequences. Shown is a summary figure of the Xenopus laevis has1 (DG42) amino acid sequence. Residues highlighted in red represent those amino acids that are conserved between all vertebrate hyaluronan synthases identified to date. To access the full sequence alignment, click here. In the full sequence alignment, colored arrows indicate the respective conserved stretches of amino acids to which degenerate PCR primers have been made and used successfully to clone partial fragments of vertebrate and invertebrate hyaluronan synthase genes.
Investigation of gene structures for the mammalian and Xenopus laevis HAS genes suggests that the hyaluronan synthase gene family evolved through a process of sequential gene duplication followed by divergence. All of the vertebrate HAS genes share the location of at least one intron (Fig. 3). Furthermore, based upon conserved gene structures and embryonic expression patterns, it appears that the Has1 gene is the mammalian equivalent or orthologue of the Xenopus DG42 gene. Accordingly, we have proposed that DG42 be redesignated Xenopus has1 (xhas1).5
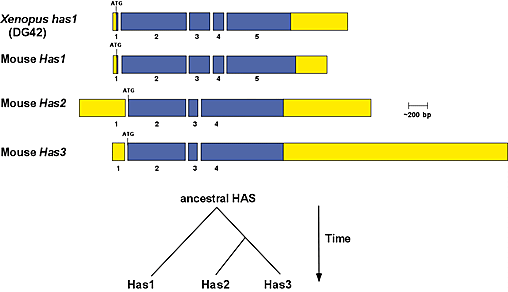
Fig. 3 Gene structure relationships of vertebrate hyaluronan synthase genes and proposed gene duplication events that gave rise to the hyaluronan synthase gene family.
Conserved exon-intron boundaries were aligned to compare the overall gene structures for the mouse Has1, Has2 and Has3 genes and the Xenopus has1 (DG42) gene. The open reading frames are shown in blue, and 5' and 3' untranslated areas are shown in yellow. The respective locations of the translation initiation codons (ATG) and stop codons (TGA, TAA) are noted. Based upon conservation of gene structure, it is proposed that the vertebrate HAS gene family arose through two sequential gene duplication events, which predate the evolution of the teleost fish. The first event duplicated an ancestral hyaluronan synthase gene to form two genes, which gave rise to the Has1 and ancestral Has2 lineages. The ancestral Has2 gene subsequently duplicated to give rise to the Has2 and Has3 genes.
The eukaryotic hyaluronan synthases are approximately 25% identical in sequence to the Streptococcus pyogenes hyaluronan synthase, spHAS. Available data suggest that it is unlikely, however, that the bacterial hyaluronan synthase3 acquired the ability to synthesize hyaluronan by horizontal gene transfer from a eukaryotic host. Based upon the location of the spHAS open-reading frame within the HAS operon and the location of related polysaccharide capsule synthase open reading frames in similar operons of other bacterial species, it is much more likely that the spHAS protein evolved from another polysaccharide capsule synthase.
As shown in Fig. 4, on the has operon, the hasA is followed by the hasB (UDP-glucose dehydrogenase) and hasC (UDP-glucose pyrophosphorylase) open reading frames. The relationship of the has operon to other polysaccharide capsule synthesis operons is shown. The cap3 operon of Streptococcus pneumoniae is required for the synthesis of a cellubiuronic acid capsule. Similar to the has operon, the cap3 operon consists of glycosyltransferase (cap3B), UDP-glucose dehydrogenase (cap3A), and UDP-glucose pyrophosphorylase (cap3C) open reading frames, although the relative positions of the glycosyltransferase and UDP-glucose dehydrogenase open reading frames are reversed with respect to their positions within the has operon. The organization of these operons is also similar to the Escherichia coli K5 capsule operon. The respective sizes (number of amino acids) of each enzyme are shown, as are the calculated amino acid sequence identities between the respective enzymes. Based upon many lines of evidence, including: (1) the location of the hasA open reading frame within the has operon; (2) the similarity of organization of the has operon to other polysaccharide capsule synthase operons; (3) the similarity in size of the hasA protein to the cap3B protein; and (4) the overall sequence identity of hasA with prokaryotic, eukaryotic, and viral glycosyltransferases, it is highly likely that hasA evolved within the context of other bacterial polysaccharide capsule synthases rather than by horizontal gene transfer from a eukaryotic host. This suggests that the evolution of the eukaryotic HAS genes and the prokaryotic HAS genes was independent, representing an example of parallel convergent evolution based upon function.
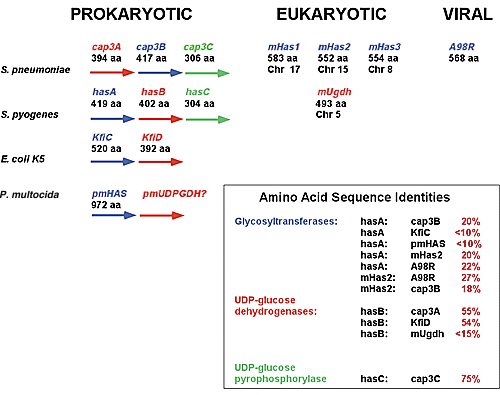
Fig. 4 Proposed evolutionary relationships between prokaryotic, eukaryotic, and viral hyaluronan synthases.
Large arrowheads indicate open reading frames within the bacterial operons. Enzymes with related activities are highlighted in the same color. Glycosyltransferases are indicated in blue, UDP-glucose dehydrogenases in red, and UDP-glucose pyrophosphorylases in green. m, mouse; Chr, chromosome. The respective chromosomes to which the mouse Has genes and the mouse UDP glucose dehydrogenase (Ugdh) genes have been mapped are shown.
Recently, the first virally encoded hyaluronan synthase, encoded by the A98R open-reading frame of the Paramecium bursaria chlorella virus, has been reported.6 This protein shares approximately 50% amino acid sequence identity with the vertebrate HAS proteins and exhibits bona fide hyaluronan synthase activity in vitro. Furthermore, algae infected by these viruses synthesize hyaluronan--the first such demonstration of hyaluronan production in the plant kingdom. The particularly high level of sequence identity between this virally encoded hyaluronan synthase and the vertebrate hyaluronan synthases suggests that this gene may indeed have been acquired by horizontal gene transfer from a eukaryotic host (Fig. 4). It appears that functional hyaluronan synthases may have evolved from two directions in prokaryotes.
The recently reported Pasteurella multocida hyaluronan synthase (pmHAS) 12 shares little sequence identity with the hasA or eukaryotic hyaluronan synthases, being more closely related to bacterial lipopolysaccharide synthase enzymes. Yet it is evident that this protein drives the synthesis of high-molecular-weight hyaluronan. It has been suggested that an open reading frame UDP-glucose dehydrogenase may lie downstream of pmHAS.12
No human or mouse genetic diseases identified to date are characterized by deficiencies in hyaluronan biosynthesis. The human HAS1 gene was localized to human chromosome (Chr) 19q13.3-13.4, and its mouse orthologue (Has1) localized to mouse Chr 17. Human HAS2 was localized to Chr 8q24.12 and its mouse orthologue (Has2) to the corresponding region of mouse Chr 15. Human HAS3 was localized to Chr 16q22.1 and its mouse orthologue (Has3) to mouse Chr 8.7
Prior to the molecular identification of the eukaryotic hyaluronan synthases, several studies investigated hyaluronan biosynthesis and the role of regulatory pathways in mammalian cell culture models. These studies were the first to demonstrate that hyaluronan biosynthesis occurs at the plasma membrane. Polymerization occurs on the inner face of the membrane, and the product is extruded or translocated through the membrane to the extracellular face of the cell.3 In fibroblast cultures, the rate of hyaluronan biosynthesis is regulated in part by cell density, and hence by the proliferative state of the cell. At low cell densities, hyaluronan biosynthesis is high, and cell motility and cell proliferation are high. At high cell densities, cell proliferation is low and hyaluronan biosynthesis is shut down.
Compounds that activate protein kinase C, such as phorbol esters, can increase hyaluronan biosynthesis in mammalian cells. Compounds that stimulate cAMP-dependent kinases also increased hyaluronan biosynthesis. Activation was largely independent of de novo protein synthesis, suggesting that pre-existing enzyme was activated. Members of the transforming growth factor beta (TGF-beta) superfamily and the platelet-derived growth factor (PDGF) family also stimulated hyaluronan biosynthesis in mammalian cell culture models. This stimulation was in part dependent upon de novo protein synthesis, suggesting that increases in hyaluronan synthase gene expression may be involved, but was also dependent upon the activation of resident proteins by protein kinase C.8 Significantly, many consensus protein kinase C phosphorylation sites are present within the predicted cytoplasmic loop of each HAS protein, suggesting that the activity of HAS proteins may be regulated in part by direct phosphorylation.
What are the functional relationships between the three vertebrate hyaluronan synthases? Expression of any one mammalian hyaluronan synthase led to hyaluronan biosynthesis by mammalian cell lines, suggesting that each functions independently as a synthase. In addition, expression of the Xenopus laevis hyaluronan synthase DG42 (xhas1) in baker's yeast led to yeast cells with plasma membrane hyaluronan synthase activity in vitro.9 Thus, a single form of HAS protein is capable of hyaluronan biosynthesis and translocation of the growing polymer. This in turn would suggest that, as predicted for the spHAS protein, the eukaryotic HAS proteins also form a pore or channel in the plasma membrane, through which the growing hyaluronan chain is extruded. Further studies will be required to elucidate the nature of this proposed pore, although work from X-ray inactivation suggests that the spHas functions as a monomer in the plasma membrane.3
Although each form of HAS protein can catalyze hyaluronan biosynthesis in eukaryotic cells, their enzymatic properties differ.c These distinct enzymatic properties may underlie physiologically distinct functions.5 Has3 is intrinsically more catalytically active than Has2, which is in turn intrinsically more active than Has1. Furthermore, analysis of hyaluronan product size generated by each HAS protein in vitro indicated that hyaluronan chain length is somehow determined by the specific HAS protein being expressed. Has1 and Has2 proteins polymerized hyaluronan chains of similar lengths (up to 2 X 106 Da), whereas Has3 polymerized much shorter hyaluronan chains, in the range of <2 X 105 Da to 3 X 105 Da. Hyaluronan chains of different lengths have different effects on cell behavior. Very short hyaluronan chains have been demonstrated to stimulate cell proliferation and to initiate signaling cascades; they may be involved in angiogenesis and inflammatory responses. High-molecular-weight hyaluronan chains have the opposite effect, inhibiting cell proliferation. Short hyaluronan chains can also be generated by degradation of extracellular hyaluronan, for instance, through the action of hyaluronidase or oxidants. The discovery that distinct HAS proteins synthesize hyaluronan chains of differing average lengths (Fig. 5) adds an additional level of regulation. The functional significance of these varying polymer lengths remains to be elucidated.
c Itano, N., Yoshida, M., Lenas, P., Yamada, Y., Spicer, A.P., McDonald, J. A., and Kimata, K., manuscript submitted for publication.
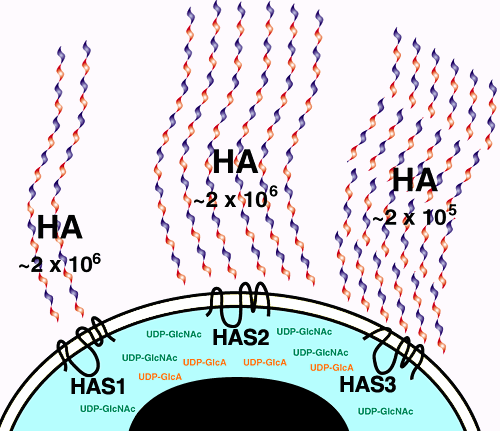
Fig. 5 Regulation of hyaluronan amount and chain length by expression of a specific HAS protein.
Biochemical characterizations of the vertebrate HAS enzymes expressed in mammalian cell culture have revealed similarities and differences between the respective mammalian hyaluronan synthase enzymes. The differences are depicted in this cartoon. Has1 produces small amounts of high-molecular-weight hyaluronan. Has2 produces significantly more high-molecular-weight hyaluronan. Has3 is the most active of the hyaluronan synthases, yet produces low-molecular-weight hyaluronan chains. The physiological significance of these differences in enzymatic activity is not yet known.
These apparent differences in enzymatic activity of the respective vertebrate HAS proteins may be related to specific amino acid residues, located within the predicted cytoplasmic loop, which vary between the respective HAS proteins, but are invariant between species ranging from zebrafish to human.5 The identification of sequences that specify rate of enzymatic activity and/or regulation of hyaluronan chain length may eventually allow us to create designer HAS enzymes, which will synthesize a given amount of hyaluronan of a certain length in a particular cell or in vitro system.
The eukaryotic hyaluronan synthases share similar degrees of amino acid sequence identities with the prokaryotic hyaluronan synthase (spHAS) and the Rhizobium NodC protein (a chito-oligosaccharide synthase). The eukaryotic HAS proteins share less identity with fungal chitin synthases. Chitin is the beta1,4 linked homopolymer of N-acetylglucosamine, best known as a constituent of fungal cell walls and of arthropod and crustacean exoskeletons. The sequence identity between Xenopus DG42 and chitin synthases led investigators to speculate on the enzymatic function of this protein. Indeed, before the molecular identification of the vertebrate hyaluronan synthase gene family, in vitro studies demonstrated that DG42 (xhas1) possessed chito-oligosaccharide synthase activity but not hyaluronan synthase activity when expressed through an in vitro transcription-translation approach. However, the reverse occurred when DG42 was expressed in baker's yeast: hyaluronan synthase activity but not chito-oligosaccharide synthase activity was demonstrated.9
Subsequently, detergent extracts of zebrafish embryos were demonstrated to possess chito-oligosaccharide synthase activity in vitro. Furthermore, this activity could be selectively removed by immunoprecipitation with a polyclonal antiserum raised against the N-terminus of the Xenopus DG42 protein.10 These data suggest, therefore, that certain members (or all members) of the hyaluronan synthase family might be able to synthesize short chito-oligosaccharides under some circumstances and hyaluronan polymers under other circumstances. Currently, it is difficult to imagine how a single HAS protein might be regulated to synthesize these related yet distinct biopolymers.
Although the existence of chitin in vertebrates is controversial, direct evidence for the presence of chito-oligosaccharides during vertebrate development has recently been demonstrated.11 Significantly, these chito-oligosaccharides could be detected only during a narrow developmental window. It is intriguing to speculate on the possible functional significance of chitin oligosaccharides in vertebrate development. To invoke a signaling role analogous to that of rhizobial nodulation factors, however, we would anticipate that a series of modifying enzymes may be required to generate the signaling competent lipochito-oligosaccharide. At this time, the possible involvement of the eukaryotic hyaluronan synthase enzymes in the biosynthesis of chito-oligosaccharides and the potential biological significance of these carbohydrate polymers in vertebrate development remain topics of controversy.
The mouse represents the model organism of choice for functional studies of mammalian genes using naturally occurring mutations, as well as transgenic and gene disruption approaches. To start to assess the respective contributions of the three hyaluronan synthases to hyaluronan biosynthesis in vivo, we have employed gene targeting in mouse embryonic stem (ES) cells to disrupt or knock out each mouse Has gene. Previously, we demonstrated that the temporal expression patterns for the three Has genes differed both in the developing embryo and in adult tissues. This, along with in vitro data demonstrating related yet distinct enzymatic activities, suggests that each HAS protein may play a highly specific role in hyaluronan biosynthesis in vivo. Has1, the least active of the three HAS proteins, drives the synthesis of high-molecular-weight hyaluronan. Thus, Has1 may play a role in maintaining a low, yet necessary level of hyaluronan biosynthesis in many cell types. Has2 is widely expressed throughout embryonic development. It is significantly more active than Has1 and also synthesizes high-molecular-weight hyaluronan. The production of large amounts of high-molecular-weight hyaluronan may have significant effects on tissue structure and volume. Thus, Has2-dependent hyaluronan biosynthesis might be predicted to play important roles in developmental processes involving tissue expansion and growth. Has3 is expressed late in embryonic development and in many adult tissues. It is the most active of the three HAS proteins, but drives the synthesis of short hyaluronan chains. Has3 expression may be activated to produce large amounts of low-molecular-weight hyaluronan, which may autonomously contribute to the pericellular matrix or may interact with cell surface hyaluronan receptors, triggering signaling cascades leading to changes in cell behavior.
Mice deficient in Has1 or Has3 activity were viable,d, e whereas mice deficient in Has2 activity were not obtained because the mutation is lethal during embryonic development. These embryos had severe developmental defects, including yolk sac and cardiac defects, and were almost entirely deficient in hyaluronan.f Thus, Has2-dependent hyaluronan biosynthesis is required for normal embryonic development, but biosynthesis involving either Has1 or Has3 is not required.
d Itano, N., Spicer, A. P., Camenisch, T., McDonald, J. A., and Kimata, K., manuscript in preparation.
e Spicer, A. P., Nguyen, T. K., and McDonald, J. A., manuscript in preparation.
f Spicer, A. P., Brehm-Gibson, T., Camenisch, T., Kubalak, S., and McDonald, J. A., manuscript in preparation.
What, therefore, are the roles for Has1 and Has3 in hyaluronan biosynthesis and function in vivo? Presently, phenotypic investigation of Has1- and Has3-deficient animals is under way. It is possible that phenotypes may arise with age, for instance, relating to joint dysfunction, other skeletal disorders, or immune-related dysfunction. In addition, there may be some degree of functional compensation built into this system. This possibility can be readily assessed by creating mice deficient in both Has1 and Has3 functions, that is, by crossing the respective mouse lines.
The identification and characterization of the eukaryotic hyaluronan synthases have revolutionized the field of hyaluronan biology. We are now able to convert essentially any mammalian cell, and possibly any eukaryotic cell, into a factory for hyaluronan biosynthesis simply by the expression of a single gene. We are also able to investigate the role of hyaluronan in developmental and disease-associated processes through in vivo genetic manipulation of mouse Has genes. Hyaluronan biosynthesis is driven by three related yet distinct enzymes in vertebrates. A combination of differential transcription of the HAS genes, intrinsic differences in enzymatic rates, and differences in resultant hyaluronan chain length provide the organism with an exquisite level of control over hyaluronan biosynthesis and function.
Acknowledgments This work was supported by individual National Research Service Award 1 F32 HL09311-01 (to A.P.S.), by American Heart Association, Arizona Affiliate, Beginning Grant-In-Aid Fellowship AZGB-19-96 (to A.P.S.), by March of Dimes Birth Defects Foundation Basil O'Connor Starter Scholar Award (to A.P.S.), by NIH grant RO1AR44689 (to J.A.M.), and by the Mayo Foundation for Medical Education and Research.