
Robert Haltiwanger
Dr. Haltiwanger received both his undergraduate degree (B.S. in Biology) in 1980 and his doctorate (Ph.D. in Biochemistry) in 1986 from Duke University. He did his doctoral work in the laboratory of Dr. Robert L. Hill on the purification and characterization of mammalian lectins, specifically the mannose receptor. He went on to do post-doctoral work with Dr. Gerald W. Hart at Johns Hopkins University School of Medicine. In Dr. Hart’s laboratory he worked on the O-linked N-acetylglucosamine (O-GlcNAc) modification. His major contribution was the purification and characterization of the enzyme responsible for addition of O-GlcNAc to proteins, the UDP-GlcNAc: polypeptide O-GlcNAc transferase. In 1992 he began his independent career as an assistant professor in the Department of Biochemistry and Cell Biology at the State University of New York at Stony Brook. There he has continued to work in unusual forms of protein O-glycosylation, including the O-fucose and O-glucose modifications. He was promoted to associate professor in 1998.
Cell surface carbohydrates were predicted to play a role in cell-cell communication and signal transduction over 30 years ago. This prediction was based on the observation that a carbohydrate-rich coat, or glycocalyx, composed of an incredibly complex array of carbohydrate structures surrounds essentially every cell in the body. The presence of such structural complexity at the cell surface, the site of cell-cell communication and signal transduction, led to the proposal that complex carbohydrates assist in the equally complex task of communication between a cell and the outside world. Nonetheless, only a few examples demonstrating a role for carbohydrates in cell-cell communication or signal transduction have been uncovered in the ensuing decades. One reason for the slow appearance of data supporting the hypothesis relates to the technical difficulty of determining oligosaccharide structures, but recent advances in analysis of oligosaccharide structures have led to a number of confirmations of the predictions made so long ago. This article describes the role of glycosylation in one particular signal transduction event, the role of O-fucose modification in the function and regulation of the Notch receptor.
The genetic locus for Notch was identified in the early part of the 20th century in one of the early genetic screens performed in Drosophila. Notch mutants show an X-linked lethal phenotype, and the locus derived its name from the fact that the females have a small notch in their wings. The gene for Notch was identified in 1985 and shown to encode a large (>300 kDal) cell surface receptor (for an excellent review on Notch, see ref. 1). Notch participates in a variety of developmental events, specifically what are known as cell fate decisions. The most detailed studies on Notch function have been performed in Drosophila where Notch functions have been shown to be necessary for the formation of numerous organs, including the neuronal system, vasculature, eyes, legs and wings. Notch homologues have been identified in all metazoans, with four in mammals (Notch1-4). Notch plays numerous roles in development in vertebrates, and defects in Notch signaling result in a variety of human diseases, including several types of cancer and developmental disorders such as Alagille syndrome, spondylocostal dystoses, and CADASIL (Cerebral Autosomal Dominant Arteriopathy with Sub-cortical Infarcts and Leukoencephalopathy). Recent work has demonstrated a link between the pathogenesis of multiple sclerosis and Notch activation. The number and magnitude of the developmental events involving Notch receptor signaling is staggering.
The Notch receptor becomes activated upon binding to its ligands, Delta and/or Serrate. Several homologues of the ligands exist in mammals (three Delta-like homologues called Delta-like1, 3 and 4; two Serrate-like homologues called Jagged 1 and 2). The ligands themselves are transmembrane proteins; thus Notch activation occurs via a ligand expressed on the cell surface of an adjacent cell (Fig. 1; for an excellent review on Notch activation see ref. 2). Notch activation proceeds through a series of proteolytic events. Ligand binding is believed to alter the conformation of the extracellular domain of Notch in such a way as to make it susceptible to cleavage by the cell-surface metalloprotease TACE (TNFα-converting enzyme). This cleavage occurs just outside of the plasma membrane, releasing the entire extracellular domain of Notch to be endocytosed by the ligand-expressing cell. Removal of the extracellular domain activates another proteolytic cleavage, catalyzed by γ-secretase, within the plasma membrane. This cleavage releases the intracellular domain of Notch as a soluble protein into the cytoplasm. The intracellular domain contains nuclear localization signals, causing it to be imported into the nucleus where is binds to members of the CSL (CBF-1/Suppressor of hairless/Lag-1) family of transcriptional regulators, resulting in activation of a number of downstream gene products. Several other cell surface receptors, including the β-amyloid precursor protein implicated in Alzheimer's disease, go through a similar series of proteolytic events, recently termed RIP, or Regulated Intramembrane Proteolysis.
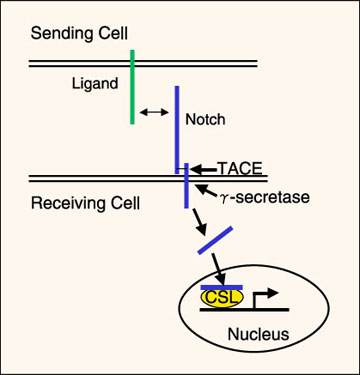
Fig. 1 Notch signaling pathway.
Notch becomes activated by binding to its ligands (Delta, Serrate/Jagged family members). Since the ligands are also transmembrane proteins, the cells must be adjacent for Notch to become activated. Ligand binding results in cleavage of the Notch extracellular domain by TACE (TNFα-converting enzyme), followed by cleavage of the intracellular domain from the membrane by γ-secretase. The Notch intracellular domain moves into the nucleus, where it interacts with members of the CSL (CBF-1/Suppressor of hairless/Lag-1) family of transcriptional regulators to activate transcription of downstream gene products. Adapted from ref. 9.
Notch activation is regulated at numerous levels3,4.For instance, expression of ligand in the same cell as Notch can exert an inhibitory effect on signaling from an adjacent cell, referred to as cell-autonomous inhibition by ligand. A number of modulators of Notch activity have been identified, including Wingless, Scabrous, Numb, Deltex and Disheveled. A novel regulator, Fringe, was initially discovered during a mutant screen for genes involved in boundary formation during wing development in Drosophila. Subsequent work showed that it functions by regulating the Notch pathway, inhibiting signaling from Serrate while potentiating signaling from Delta. Interestingly, Fringe's effects are cell autonomous with respect to Notch (i.e. Fringe needs to be expressed in the same cell as Notch to function) and Fringe is a secreted protein. These observations led to the suggestion that Fringe may mediate its effects by either binding to or post-translationally modifying the extracellular domain of Notch. The fact that Fringe showed weak sequence similarity to several bacterial glycosyltransferases raised the possibility that Fringe functions by altering the structure of the carbohydrate chains on Notch.
The extracellular domain of Notch is modified with several forms of glycosylation including O-fucose and O-glucose5. These unusual forms of O-linked glycosylation occur on the hydroxyl groups of serine or threonine residues at consensus sequences within epidermal growth factor (EGF)-like repeats6. An EGF-like repeat is a protein motif defined by the presence of six conserved cysteine residues forming three disulfide bonds, and the consensus sequences for O-fucose and O-glucose are found in the context of these cysteines. For instance, the O-glucose site occurs between the first and second conserved cysteine of an EGF-like repeat at the sequence C1XSXPC2, while the O-fucose site is found between the second and third conserved cysteines at the sequence C2X4-5S/TC3 (Fig. 2). The Notch extracellular domain is composed of up to 36 tandem EGF-like repeats (e.g. mammalian Notch1 and Notch2, Drosophila Notch), many of which contain consensus sequences for O-fucose and O-glucose saccharides (Fig. 3). The locations of many of the sites for glycosylation are evolutionarily conserved, suggesting that the sugars play an important role in the biology of Notch.
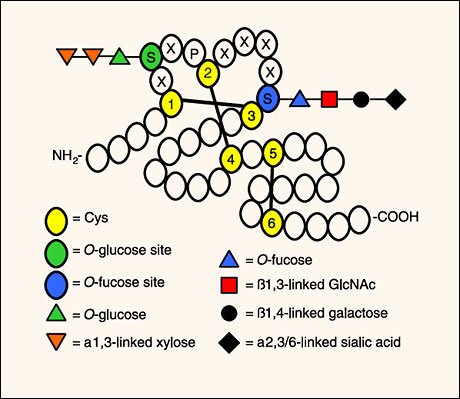
Fig. 2 Some EGF-like repeats are modified with O-fucose and/or O-glucose saccharides.
A schematic representation of an EGF-like repeat is shown, highlighting the cysteines involved in internal disulfide bonds (black lines) and the putative consensus sequences for addition of O-fucose and O-glucose saccharides. Representations of the potential O-fucose and O-glucose structures are shown, although variability of the structures is seen on different EGF-like repeats. For instance, EGF-like repeat 1 from human factor VII has a simple O-fucose modification, whereas EGF-like repeat 1 from human factor IX has the complete tetrasaccharide6.
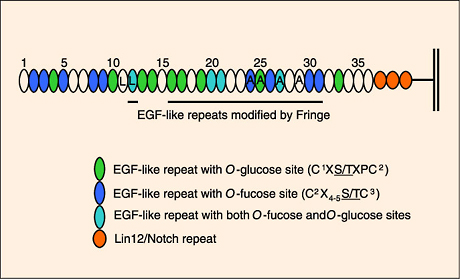
Fig. 3 Many of the EGF-like repeats in the extracellular domain of Notch contain sites for O-fucose and/or O-glucose modifications.
Only sites conserved in Notch1 and Notch2(human and mouse) as well as Drosophila Notch are shown. In addition, O-fucose bearing EGF-like repeats that occur in regions of the molecule known to be modified by one of the Fringe proteins are underlined. "L" denotes EGF-like repeats involved in binding to ligands. "A" denotes EGF-like repeats to which Abruptex mutations map. Adapted from ref.12.
Although little is known about the O-glucose modifications, a number of studies suggest that the O-fucose modifications are essential for Notch function. Reduction in expression levels of protein O-fucosyltransferase I (O-FucT-1), the enzyme responsible for addition of O-fucose to EGF-like repeats, by RNA interference in Drosophila results in Notch-like phenotypes7. Detailed studies have demonstrated that reduction of O-FucT-1 levels affects Notch function in many different developmental contexts, suggesting O-fucose is essential for all aspects of Notch signaling. Genetic ablation of O-FucT-1 in mice results in a phenotype more severe than any single Notch knockout, suggesting O-fucose modifications are essential for all Notch isoforms8. In addition, cells with defects in GDP-fucose synthesis (the fucose-donor for all fucosyltransferases) do not support Notch signaling9,10. These results strongly suggest that Notch must be modified with O-fucose to function.
In addition to being essential for function, O-fucose plays a role in modulation of Notch activity. Recent studies have demonstrated that the Fringe family of proteins are O-fucose specific β1,3-N-acetylglucosaminyltransferases (β1,3GlcNAcT)9,11. In the absence of Fringe, Notch is modified with O-fucose monosaccharides. In the presence of Fringe, O-fucose on several (but not all) of the EGF-like repeats is modified with a ゚1,3-linked GlcNAc12. This disaccharide can be further elongated with a galactose and sialic acid to make a mature tetrasaccharide (Sia-α2,3/6-Gal-β1,4-GlcNAc-β1,3-Fuc-α1-O-Ser/Thr, Fig. 2). Fringe alters the ability of Notch to respond to its ligands by changing the structure of O-fucose on Notch: inhibiting signaling from Serrate/Jagged but potentiating signaling from Delta. Addition of the sialic acid does not appear to be essential for inhibition of signaling from Jagged1, suggesting that the trisaccharide Galβ1,4-GlcNAcβ1,3-Fuc is the minimal structure necessary for Notch activity to be altered10. Fringe provides a specific example of how the activity of a signaling receptor can be regulated by alterations in its glycosylation state (for recent reviews see ref. 13,14).
Although it is now clear that the O-fucose modifications on Notch are essential for its function and for Fringe to modulate its activity, the molecular mechanisms for how these saccharides affect Notch are not well understood13,14. It is not known why O-fucose is required for Notch function. Preliminary studies show that Notch is expressed at the cell surface in the absence of O-fucosylation suggesting that O-fucose is not required for proper cell-surface expression7. Thus, the sugars could participate directly in binding to the ligands, or they may play a role in causing Notch to assume a certain conformation necessary for ligand binding and/or proteolysis. In addition, the mechanisms by which the Fringe-mediated alterations in O-fucose structures modulate Notch signaling are not known. One complication for all of the available models is that Fringe has opposite effects on signaling from the two classes of ligands, potentiating signaling from Delta ligands while inhibiting signaling from Serrate/Jagged ligands. This suggests that at least two mechanisms may be at play. The change in O-fucose structure could have a direct effect on ligand binding, or it may induce a conformational change in Notch that indirectly alters ligand binding or susceptibility to TACE. Alternatively, another protein may be involved, and the alterations in O-fucose structure could affect the ability of Notch to interact with this protein. Several proteins are known to interact with the EGF-like repeats of Notch and affect signaling (e.g. Scabrous, Wingless). In addition, ligand expressed in the same cell as Notch can exert an inhibitory effect. Alterations in the interactions of Notch with any of these proteins, mediated by a change in O-fucose structure, could provide a mechanism for Fringe action. Interestingly, O-fucose modifies EGF-like repeats involved in ligand binding and in the Abruptex region (Fig. 3). Abruptex mutations result in hyperactivatible forms of Notch; thus the region in which they occur is believed to be inhibitory. The presence of O-fucose saccharides at these sites may offer some clues to the function of these sugars. More work needs to be done to determine which of these mechanisms is at play with each of the ligands.
In addition to not knowing the molecular mechanisms of how the O-fucose glycans affect Notch function, little is known about the function of the O-glucose glycans on Notch. Notch is modified with O-glucose in both the monosaccharide and trisaccharide (proposed structure: Xyl-α1,3-Xyl-α1,3-Glc-β1-O-Ser/Thr) forms5. The sites of O-glucose on Notch have not yet been mapped, but preliminary data from our laboratory indicates that the predicted sites are in fact modified. The O-glucose sites on Notch are as highly conserved across evolution as the O-fucose sites (Fig. 3), suggesting an equally important role for these modifications in Notch biology. Enzymatic activities have been identified for the protein O-glucosyltransferase responsible for addition of O-glucose to EGF-like repeats15 and for the two predicted xylosyltransferases16,17, but a great deal of work remains to identify these enzymes and their biological function.
The proposal that cell surface glycans participate in cellular communication is being realized. The past decade has seen numerous examples of specific biological events being controlled or regulated by cell surface glycoconjugates. The involvement of sialyl Lewis x structures in recruitment of leukocytes to sites of inflammation, the role of heparin sulfate proteoglycans in Wnt, hedgehog and TGFβ signaling pathways, the role of O-mannose saccharides in binding of α-dystroglycan to the extracellular matrix, and the modulation of NCAM interactions by alterations in polysialic acid are all clear examples of the role cell surface glycans play in modulating significant biological events. The role of O-fucose on Notch is a new addition to this ever-growing list. The discovery that Fringe is a glycosyltransferase resulted directly from the ability to compare genomes from a variety of species. The next decade should be an exciting time, as more putative glycosyltransferases are shown to synthesize sugar structures essential for a variety of biological functions.